The fight against one of the nation’s leading diseases has just gotten a lot more personal. Diabetes
mellitus is a chronic disease that compromises the body’s ability to regulate blood glucose, leading to
a variety of complications that may worsen over time. Each person has a diverse health background,
so every person’s diabetic diagnosis falls within a broad range of conditions. Diabetes is not merely a
single-faceted disease but rather is a broad spectrum of disease subtypes or less common versions
of the disease. This spectrum of disease conditions in diabetes could contain unique diabetic cases
specific to each person’s genetic code and this poses a major problem for current treatment methods
which provide more general therapies that may not be optimal for every individual. While still in
infancy, personalized medicine is coming into the spotlight as a next generation cure alongside a
shift in diabetic research towards the genomic level–the most individualized approach one can take–
to deliver targeted therapies.
Dr. Amit Majithia, an Assistant Professor of Endocrinology at the Department of Medicine and Pediatrics at the UC San Diego School of Medicine, researches novel genetic methods of detection and treatment for insulin resistance.¹ Currently, he directs his efforts towards providing a more personalized therapeutic approach for type 2 diabetic patients. The Majithia lab uses a multi-disciplinary approach that combines computational bioinformatics and large-scale genomic techniques in order to further understand and classify genetic variants, or different versions of the same gene that vary in nucleotide sequence. The Majithia lab is specifically looking for genetic variants that are linked to an increase in insulin sensitivity by analyzing the functions of the genes and matching them to specific phenotypes through a parallel-line bioassay technique¹ which is an experimental technique that measures a target substance in various dilutions.² Through the functional characterizations of the missense genomic variants, a more efficient understanding of their impact on the function of genes related to diabetes can be made in order to provide deeper insights into diabetic treatment.
Currently, diabetic patients use traditional testing methods such as blood tests on a case-by-case basis when testing for the presence of potential, diabetically-linked variants. Rather than pursuing this
old-fashioned route of testing, Dr. Majithia and his team have turned to the construction of a “lookup table” in which every possible mutation is synthesized and tested before it is discovered in a person. This is accomplished by using synthetic biological techniques and parallel cellular assays that are centered around identifying causes and susceptibilities of disease. The main genetic variants targeted are those that are hypothesized to correspond to an increase in insulin sensitivity through missense variations that produce proteins with altered functions.¹ Through this new, proposed research method, a patient who is diagnosed as either pre-diabetic or diabetic can have their DNA sample taken for whole genome sequencing. Unique genetic variants that are linked to increased insulin sensitivity¹ are identified from the patient’s DNA sample, and functional characterization of these missense variants is done in order to determine the best course of treatment. As further mutations are discovered via patient samples, the “lookup table” enables researchers to cross-check previously unknown genetic variants in patients to synthetically tested variants, thereby quickening the timeline for clinical testing and treatment.
One of the genes the Majithia lab studied, peroxisome proliferator-activated receptor gamma (PPARG), contained certain mutations which were linked to an increased risk of type 2 diabetes and lipodystrophy, the abnormal loss and distribution of fat.³ PPARG is one of many genes that were functionally classified in the Majithia lab in an effort to characterize and encode a table of all 9,595 single-amino acid substitutions found in peroxisome proliferator-activated receptor gamma (PPARγ), which is a nuclear receptor that is encoded by the PPARG gene.³ Under Dr. Majithia’s lab, fifty-seven other missense variants of PPARG were analyzed, out of which six variants were classified as pathogenic, or related to disease.³ In other words, these six particular variants were correlated with a higher probability of susceptibility to lipodystrophy and type 2 diabetes. With the sequencing of more protein encoding sequences like PPARG, more missense variants are on track to be identified. The characterization of novel variants will ultimately provide researchers with a more comprehensive understanding of specific genes that are correlated with diabetes.
Researchers at Dr. Majithia’s lab realized that current treatment and diagnostic techniques had the potential to provide a baseline for upgraded treatment and diagnostic technologies as they looked into the current mechanisms of continuous glucose monitoring (CGM) sensors. CGM readings are primarily used before and after meals in order to inform patients of their glucose levels. These readings allow patients to plan meals and insulin intake accordingly. Dr. Majithia noticed that current diabetic devices such as insulin pumps provide patients with constant information on blood glucose levels which could allow patients to increase their capacity to manage their diabetes. However, the standard CGM methods do not provide the most reliable and up-to-date results. Following this line of thought, the Majithia lab repurposed CGM sensors to provide further, reliable readings with consistent analysis of blood glucose
levels that are now used in technologies for type 1 diabetic patients. This is done by improving the accuracy of CGM readings by showing a wider range of blood glucose readings that presents less restricted rate of change arrows. Many current CGM devices’ rate of change arrows are utilized for convenience as they present rounded off number readings, and through repurposing these arrows, they can provide much more reliable information for patients to determine their insulin dosages.
For type 2 diabetes, traditional testing currently involves a blood sampling technique that requires patients to prick their fingers throughout the day to obtain blood glucose readings. Similarly to the continuous monitoring technology in an EKG, a new method of diabetes monitoring for type 2 diabetic patients could potentially be developed that adopts the CGM method, in which patients monitor their glucose levels before and after meals. Instead of multiple finger pricks throughout the day, a single finger prick is sufficient for this newly improved blood test. Finger prick blood testing uses test strips that contain the enzyme glucose oxidase, which reacts with a patient’s blood. The reaction between blood and glucose oxidase generates an electric impulse that is measured by the blood glucose monitoring device; the more glucose there is in the blood, the greater the magnitude of the current (reflected as a numerical value in the device). This blood glucose level number is taken together with measurements of blood pressure and cholesterol levels to yield a comprehensive reading that determines how much insulin a patient should inject daily. Thus, in utilizing an improved CGM method that is traditionally found in type 1 diabetic patients’ technologies, type 2 diabetic patients may also be able to receive accurate, up-to-date readings in regards to their blood glucose levels and therefore be able to make better informed decisions in their everyday insulin treatment. It should be noted that treatment for type 2 diabetic patients does not focus solely on insulin administration but also can involve the use of CGMs to help patients make informed, lifestyle decisions.

Diabetes is a unique disease that can be affected by a person’s genetic makeup and influenced by their lifestyle and environmental factors. As such, an accurate diagnosis of diabetes goes beyond the general categorizations of Type 1 and Type 2.
There are also novel analytic methods being researched that focus on patterns hidden within blood glucose monitor data rather than on individual blood glucose values. Diagnostics and testing go hand in hand in the Majithia lab, where efforts are focused on developing more accurate and precise diagnostic tools that can identify more diabetic subtypes. Glucotypes are measurements of glucose variability patterns taken from comparisons of input from CGM systems and diabetic diagnostic measurements, such as daily insulin secretions and levels of insulin resistance. These glucotypes are used to formulate meaningful diagnoses of specific type 2 diabetes subtypes and represent a form of technology that could provide each patient with a diagnosis of a specific diabetic subtype rather than a generic diagnosis of type 2. The Majithia lab analyzes genetically-linked and pharmacologically sourced blood glucose data from both non-diabetic and diabetic patients and uses algorithms to generate more personalized and accurate glucotypes for each individual. Subtyping also has the potential to streamline categorization of the complications that correlate with different diabetic subtypes by providing further specificity about the type of diabetes and its symptoms. Both of these developments will further clarify the understanding of diabetes and generate more personalized methods of treatment.
Genetic testing is another practice that can potentially provide personalized diagnoses of diabetes, thereby helping to illuminate once indistinguishable diabetic types for patients. Through previously mentioned methods, such as the Majithia lab’s construction of a genomic ‘lookup table’, an individual may receive a more specific diagnosis of the subtype of diabetes they have. In specific diabetic cases that produce rare genetic diagnoses such as monogenic diabetes or diabetes caused by a single point mutation, genetic testing can be used to categorize what subtypeâ· of diabetes the individual has. In cases concerning newborn and young children, new genetic technologies are being employed to provide more accurate diabetic diagnoses such as the diabetic subtype maturity-onset diabetes of the young (MODY).â· Oftentimes, patients with MODY are misdiagnosed with type 1 diabetes and placed on standard insulin treatments which are not effective courses of therapy for MODY. Genetic testing, however, is more likely to help elucidate a patient’s specific condition, allowing patients to successfully switch from an insulin based treatment to a new class of drugs like sulfonylurea (SU), which are better suited to alleviate the complications of MODY. Personalized diagnosis through genetic testing is the first of many steps that will enable doctors to provide patients with more reliable descriptions of their specific forms of diabetes.
Personalized treatment for diabetes has already begun to take root in the pharmacogenomic sector. Pharmacogenomics is the study of how the genetic makeup of an individual reacts to certain drugs.â· Every individual’s genetic makeup is different, and thus each person will generate slightly different responses towards each drug. Pioneering work on more personalized pharmacogenomics has been aided by current research on genetic variants of diabetes, which take the form of genomic libraries. These massive tables of data, such as the PPARG table that Dr. Majithia and his colleagues created, provide researchers with comprehensive genotypic and phenotypic data that are useful for the detection of treatment response patterns and drug responsiveness in diverse groups of patients.â· With more accurate genomic data that help tailor treatment for the individual, there could be decreased risks of adverse drug-related events, leading to more optimal treatment.
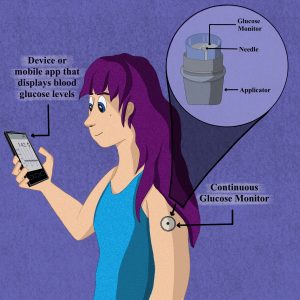
Modern diabetic technologies use mobile apps that synchronize with continuous glucose monitoring patches to monitor blood glucose levels throughout the day.
Personalized therapy is still in its early stages, and there is much more research and testing that needs to be done in order to implement it clinically. Despite the research that still needs to be conducted, the efforts of Dr. Majithia and his lab have advanced the progress towards technological and genomic advancement of personalized medicine for diabetes. With the proper biotechniques, diabetes research rooted in personalized medicine could establish a model for how to approach other diseases. Current treatments for diabetes are still far too generalized to provide optimized treatment for each individual. In taking a genetic, more personalized route, however, better treatment methods and more accurate diagnoses can be provided to patients. The future is personal and there is so much more to explore about the individuality of each body.
REFERENCES
- Research | Majithia Lab | UC San Diego School of Medicine. medschoolucsdedu. [accessed 2021 Apr 21]. https://medschool.ucsd.edu/som/medicine/divisions/endocrinology/research/labs/majithia/research/Pages/default.aspx.
- Stegmann DR. 2021 Feb 9. Parallel-line potency assays. PLA 30 — Software for Biostatistical Analysis. [accessed 2021 Apr 21]. https://www.bioassay.de/analytical-methods/experiments/parallel-line-potency-assays/.
- Majithia AR, Tsuda B, Agostini M, Gnanapradeepan K, Rice R, Peloso G, Patel KA, Zhang X, Broekema MF, Patterson N, et al. 2016. Prospective functional classification of all possible missense variants in PPARG. Nature Genetics. 48(12):1570—1575. doi:10.1038/ng.3700.
- Majithia AR, Wiltschko AB, Zheng H, Walford GA, Nathan DM. 2017. Rate of Change of Premeal Glucose Measured by Continuous Glucose Monitoring Predicts Postmeal Glycemic Excursions in Patients With Type 1 Diabetes: Implications for Therapy. Journal of Diabetes Science and Technology. 12(1):76—82. doi:10.1177/1932296817725756.
- MIT School of Engineering |» How do glucometers work? 2011. Mit Engineering. https://engineering.mit.edu/engage/ask-an-engineer/how-do-glucometers-work/.
- What Is Glucotyping? New Method of Detecting Glucose Dysregulation. 2018 Aug 18. Diabetes In Control A free weekly diabetes newsletter for Medical Professionals. [accessed 2021 Apr 21]. http://www.diabetesincontrol.com/what-is-glucotyping/
- Xie F, Chan JC, Ma RC. 2018. Precision medicine in diabetes prevention, classification and management. Journal of Diabetes Investigation. 9(5):998—1015. doi:10.1111/jdi.12830.
WRITTEN BY MARCELLA KU
Marcella is a General Biology Major from Roger Revelle College. She will be graduating in 2023.
FROM SALTMAN QUARTERLY VOL. 18
To read the original version, please click here. To read the full version on our website, please click here. To read more individual articles, please click here.