Analyzing recent developments for infectious disease treatments and technologies for malaria and tuberculosis
For as long as people have existed, there have been diseases and treatments. Historically, viruses that have caused cold-like symptoms were deemed serious medical risks, but such diseases that ravaged communities were eventually eradicated by modern treatment techniques such as vaccines. Vaccines, antibiotic treatment for bacterial-caused diseases, and even common practices such as hand-washing were all medical breakthroughs that contributed to the prevention of illnesses throughout history.
In light of the COVID-19 pandemic, breakthrough medical strategies and technologies have been receiving more public attention and praise, such as: automated disinfection protocols, rapid viral load testing, and RNA-based vaccines. While technologies such as these are being developed more rapidly and efficiently than ever, the same cannot be said for treatment distribution. Despite the quick production of COVID-19 vaccines, many countries have struggled to obtain adequate supplies to protect their populations because other countries over-buy supplies or deliberately reduce access. Unfortunately, the same issue arises for the treatments of two other dominant global health threats: tuberculosis and malaria.
Unlike their more manageable infectious counterparts, such as the common cold, malaria and tuberculosis have been prevalent diseases for thousands of years and continue to predominantly affect under-resourced countries. These two diseases, caused by parasites and bacteria, respectively, have medical professionals and scientists working tirelessly to eradicate them. But what new, long lasting treatment options are being devised to fight tuberculosis and malaria with higher precision and accuracy? Such issues are being researched today at UC San Diego’s Winzeler lab, which delves into malaria drug efficiency, and the Sailor lab, which analyzes nanoparticle drug delivery for tuberculosis treatment.
BODY CELLS AND PARASITES
Despite being around for thousands of years, treatments for malaria remain inefficient and somewhat convoluted. Unlike many familiar diseases like the flu, malaria is not caused by a virus but instead by the unicellular Plasmodium parasite, which is transmitted to humans through bites from infected mosquitoes. Inside a host, these parasites enter the liver, and once mature, they will leave and enter the patient’s bloodstream and infect red blood cells, thus causing anemia, fatigue, and a host of other critical symptoms. Since the disease is parasitic rather than bacterial, there is no set treatment that can kill the parasites while ensuring that healthy human cells remain intact. This is because malaria can proliferate either through the bloodstream or through the liver pathway, making it difficult to provide pathway-specific treatment. Because of this, there is a risk of the parasites developing resistance to common treatments, similar to how bacteria are often able to develop resistance to frequently-used antibiotics as analyzed by Kokwaro and colleagues in a 2009 study. The imminent threat of parasitic resistance combined with the need to kill as many parasites as possible results in malaria treatments using combinations of several drugs rather than a singular medication.
At UC San Diego, the Winzeler Lab is searching for a more effective treatment for malaria. Historically, it has been challenging to develop quality treatment that ensures the protection of healthy blood cells while also killing parasites in infected blood cells. For example, a popular drug commonly known as Malarone can lead to liver failure while removing parasites in liver cells. At the Winzeler Lab, third-year undergraduate student Sindhu Daggupatti and her colleagues are focusing their research efforts on the Malaria Drug Accelerator (MaLDA) project. MaLDA is a nationwide project in which 15 malaria-focused labs research efficient malaria drugs that minimize bodily damage and effectively target phenotypic markers.
One of the key elements of malaria drug development is recognizing the Plasmodium parasite’s characteristics, with particular emphasis on areas within the body containing high quantities of Plasmodium. As Plasmodium are primarily found in the liver and blood, the health of these two areas is a key indicator of the effectiveness of possible treatments. If a treatment can kill the most Plasmodium while keeping most body’s own cells intact, there is a high probability that the medication would be more efficient and beneficial for long-term use than its competitors. On a typical day, Daggupatti analyzes the effect of diluted medications on samples of parasitically-infected liver cells that are then scanned to determine whether the drug is able to not only kill parasites but also keep the human body cell intact. A similar test is performed on blood samples containing Plasmodium under similar standards as the liver assays.
The steps taken by the Winzeler lab and others like them are the critical first step to developing a more efficient treatment for this disease. Daggupatti’s efforts within the groundbreaking MaLDA project enabled further testing on drug efficacy and improved predictions of parasitic gene resistance to drugs. Such efforts are paving the way for potentially new indicative properties of malaria for new targets of drug efficiency.
SMALLER IS BIGGER
While drug resistance is one of the leading issues that infectious disease treatment and technology faces, novel infectious disease labs at UC San Diego have been looking into alternative routes through the use of similar drugs transported to target body locations via targeted mechanisms. Drug resistance is not a challenge faced exclusively in the response to treatments against parasitic diseases; in fact, resistance is most commonly associated with bacteria. Scientists have begun to look into mechanistic alternatives for bacterial diseases such as tuberculosis that can allow for common treatment drugs to be used.
Tuberculosis, one of the globe’s dominant infectious diseases caused by the bacteria Mycobacterium tuberculosis, is an airborne respiratory illness that requires long-term and consistent treatment. Tuberculosis is devastating for those who contract it and has the greatest impact on the least-resourced nations. The slew of medications and doctor’s appointments required to stave off the bacterial infection are often significant socioeconomic barriers for many impacted communities. One of the most pervasive issues faced when treating tuberculosis is the proportion of antibiotic-resistant bacterial lesions – colonies of bacteria on the lungs with genetic immunity to typical antibodies. This can either be a result of a random mutation occurring in a colony of bacteria or developed over time through consistent exposure to the same antibiotics. When treating tuberculosis, several types of antibiotics are used, and each exposure to a new drug only increases the chances that the disease-causing bacteria will develop immunity.
 The Sailor lab at UC San Diego opens a new realm of possibilities for advanced disease treatments through nanoparticle research with the goal of precisely delivering medication to reduce the likelihood of genetic resistance developing. Third-year undergraduate student Rachel Lee from UC San Diego, a researcher at the Sailor lab, and her lab partner Rachel Myers, a UCSD Materials Research Science Engineering Center Research Experiences for Undergraduates summer intern from University of Maryland Baltimore County, have worked on nanoparticle technology to deliver a commonly used antibiotic drug for tuberculosis, called rifampicin, in a more targeted manner into tuberculosis bacteria. This targeted treatment allows for direct loading antibiotics into bacterial lesions in the lungs. With a toxic drug like rifampicin, this nanoparticle technique is much more efficient than unspecifically loading the bloodstream. Rather than exposing and impacting the cells of the whole body and other possible bacterial populations, the medication is only exposed to the specific bacteria it is trying to kill, reducing the likelihood of antibiotic resistance and lowering the risk of harmful side effects.
The Sailor lab at UC San Diego opens a new realm of possibilities for advanced disease treatments through nanoparticle research with the goal of precisely delivering medication to reduce the likelihood of genetic resistance developing. Third-year undergraduate student Rachel Lee from UC San Diego, a researcher at the Sailor lab, and her lab partner Rachel Myers, a UCSD Materials Research Science Engineering Center Research Experiences for Undergraduates summer intern from University of Maryland Baltimore County, have worked on nanoparticle technology to deliver a commonly used antibiotic drug for tuberculosis, called rifampicin, in a more targeted manner into tuberculosis bacteria. This targeted treatment allows for direct loading antibiotics into bacterial lesions in the lungs. With a toxic drug like rifampicin, this nanoparticle technique is much more efficient than unspecifically loading the bloodstream. Rather than exposing and impacting the cells of the whole body and other possible bacterial populations, the medication is only exposed to the specific bacteria it is trying to kill, reducing the likelihood of antibiotic resistance and lowering the risk of harmful side effects.
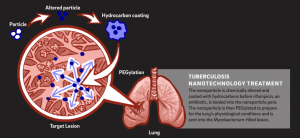
 In their experiment, a nanoparticle, which is a hollow sphere 100,000 times smaller than the diameter of a piece of hair, is utilized for a variety of chemical and biological reactions. It is first chemically altered and then loaded with the rifampicin drug to target tuberculosis-infected lesions. The nanoparticle pore is coated with hydrocarbons to allow for the antibiotic rifampicin to be placed inside and then PEGylated. PEGylation is a chemical process that synthesizes polymers onto the nanoparticle and increases drug stability, reducing the amount of required doses of medication.This sequence of chemical processes stabilizes the drug and ensures that the process and the medication will perform properly in human cells. During the rifampicin loading process, calcium chloride is added to reduce the time it takes to add the drug into the nanoparticle from 24 hours to around 12 hours. This multi-step technique of drug loading led to observed significant mass loading of rifampicin into the nanoparticle. Through the usage of this nanoparticle, the rifampicin antibiotic is able to be delivered to the tuberculosis-infected lesions more effectively as the drug is stabilized and the targeting route is more specific.
In their experiment, a nanoparticle, which is a hollow sphere 100,000 times smaller than the diameter of a piece of hair, is utilized for a variety of chemical and biological reactions. It is first chemically altered and then loaded with the rifampicin drug to target tuberculosis-infected lesions. The nanoparticle pore is coated with hydrocarbons to allow for the antibiotic rifampicin to be placed inside and then PEGylated. PEGylation is a chemical process that synthesizes polymers onto the nanoparticle and increases drug stability, reducing the amount of required doses of medication.This sequence of chemical processes stabilizes the drug and ensures that the process and the medication will perform properly in human cells. During the rifampicin loading process, calcium chloride is added to reduce the time it takes to add the drug into the nanoparticle from 24 hours to around 12 hours. This multi-step technique of drug loading led to observed significant mass loading of rifampicin into the nanoparticle. Through the usage of this nanoparticle, the rifampicin antibiotic is able to be delivered to the tuberculosis-infected lesions more effectively as the drug is stabilized and the targeting route is more specific.
This research project has inspired Lee and Myers to continue to pursue this mechanism of treatment for future infectious diseases as well. In the near future, they aim to characterize each step of the drug-loading process further in hopes of maximizing final rifampicin concentration in each nanoparticle.
A NEW PROMISE
Such novel techniques, like these more effective drug therapies, illustrate many possible paths for future medications to reduce the risks of drug resistance and increase overall effectiveness. Costs could be lowered for research, development, and distribution of these life-saving treatments in the long term. For patients in medically underserved areas or those unable to consistently attend treatments, this kind of updated treatment regimen would drastically improve their ability to recover through equitable cost and distribution of these drugs. As these technologies become more refined, there can be higher cost-efficiency in drug synthesis, leading to affordable treatment and medication for less socioeconomically developed nations.
Recent developments in infectious disease treatment via alternative drug mechanism routes and new drug synthesis aid in providing new routes for effective, simplified treatment and serve as progress towards global, equitable treatment and technologies. Ranging from malaria drug synthesis to nanoparticle rifampicin delivery for tuberculosis, these promising therapeutics are on the forefront of new disease research that can help change how we treat dozens of bacterial and viral diseases or even change how medications are developed. In time, these research advancements could greatly reduce infectious diseases, which currently possess great power over many nations, as our technology becomes more advanced, and our people even stronger.
Written by Marcella Ku & Emma Svendsen
Marcella is a third year student double majoring in General Biology and Global Health.
Emma is a first year student majoring in General Biology.

