EXTRACELLULAR ELECTRON TRANSFER AND MICROBIAL NATURAL PRODUCTS
Oxygen is an essential part of the electron transport chain, functioning as the terminal electron acceptor in aerobic respiration.¹ Without oxygen, an aerobic cell is unable to complete the final step of electron transfer, causing the transfer process to halt. When this occurs, the intermembrane proton gradient is not maintained, and ATP synthase is unable to generate ATP for the cell.¹ The resulting lack of oxygen inhibits necessary cellular pathways and the cell eventually dies due to asphyxiation. However, some bacteria have developed ingenious ways of circumventing this issue. Apart from more traditional forms of low-oxygen respiration, such as the use of high-affinity terminal oxidases by microaerophilic bacteria, several groups of bacteria have evolved unique means of respiration that utilize oxygen indirectly.² This indirect process, known as extracellular electron transfer, can occur through a variety of different methods. Some methods include external longitudinal electron transfer by cable bacteria (over centimeter scale distances), conductive nanowires by dissimilatory metal reducing bacteria, and the use of exogenous humic substances by various soil and marine sediment bacteria.³,, However, one recently discovered type of extracellular electron transfer is performed by endogenously-produced redox-active natural products.
Microbial natural products (also known as secondary metabolites) are organic molecules produced outside of a microorganism’s primary metabolism. Natural products have various roles, acting as anything from signaling molecules to antibiotics. They have also been a fruitful source for modern drug discovery.â· However, one group of natural products, small-molecule extracellular electron shuttles (SMEES), blurs the line between primary and secondary metabolites. While most natural products do not contribute directly to growth or metabolism, SMEES have been shown to directly participate in cellular respiration and survival in oxygen limited environments.⸠While several classes of SMEES have been identified, phenazine secondary metabolites are the best characterized endogenously produced extracellular electron shuttles (Figure 1).â¸,â¹ Phenazine metabolites, especially those produced by the pathogenic bacteria Pseudomonas aeruginosa, have been shown to transfer electrons from oxygen deficient areas (such as in the center of bacterial biofilms), to areas with high oxygen (such as the periphery of biofilms) in order to perpetuate the electron transport chain when oxygen is limiting.â¹
MECHANISM OF PHENAZINE MEDIATED EXTRACELLULAR ELECTRON TRANSFER
Phenazine natural products are an anomaly in the world of microbial chemical ecology. While most characterized bacterial natural products have yet to be linked to their ecological roles, phenazines have been tied to several.¹â° Phenazines are known to function as virulence factors, participate in biofilm formation, late-stage quorum sensing, extracellular metal reduction, and antibacterial activity.¹â° However, their ability to shuttle electrons out of the cell under oxygen limiting conditions while stabilizing the intracellular redox state is the most unique aspect of these molecules.¹¹ While scientists first reported in 1931 that Pseudomonas aeruginosa produced an unidentified diffusible, redox active substance in low-oxygen conditions, a mechanism for phenazine-mediated extracellular electron transfer (PMEET) was not proposed until late 2020.â¹,¹²
PMEET has been best studied in Pseudomonas aeruginosa biofilms, where phenazines stimulate bacterial swarming motility and contribute directly to colony morphology.¹³ Once a biofilm has been formed, cells deep within the biofilm are quickly deprived of oxygen due to normal metabolic activity. In an article by Saunders, et al. two multi-step mechanisms for PMEET have been proposed (Figure 2).â¹ Analogous to the intracellular electron transport chain, the steps involve transferring electrons from molecules with a low redox potential to those with a higher redox potential. Several phenazine metabolites–phenazine-1 carboxamide (PCN), phenazine-1 carboxylic acid (PCA), and pyocyanin (PYO), listed in order of increasing redox potential–act in concert to transfer electrons from the interior of the biofilm to the periphery of the biofilm where oxygen is available to be the final acceptor of electrons.â¹
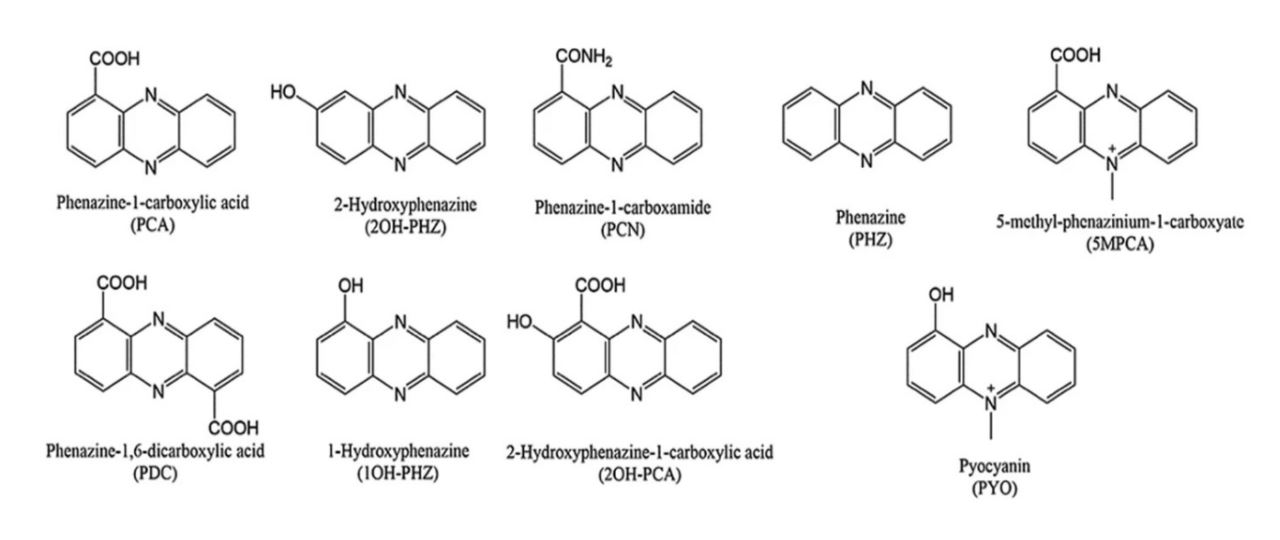
The core phenazine moiety, as shown in the bottom left by PHZ, is the core of all phenazine metabolites. The various functional groups observed in each molecule directly affect the characteristics of the metabolite, such as hydrophobicity and redox potential. Bilal (2017) Microbial Biotechnology.â¸
The two proposed mechanisms both begin with a low-potential phenazine (either PCN or PCA) removing electrons from an oxygen-deprived cell by a currently unknown mechanism.â¹ The next step involves reduced PCN or PCA molecules freely diffusing throughout the biofilm until they reach the periphery.â¹ Then, reduced low-potential phenazines transfer electrons onto PYO, a higher potential phenazine metabolite.â¹ Pyocyanin is preferentially retained at the periphery of the biofilm due to intercalation with extracellular DNA (eDNA), a major component of biofilms.¹ From there, PYO transfers electrons to molecular oxygen, forming water.â¹ Pyocyanin then binds with eDNA and the cycle starts again. The second proposed method follows a similar pattern as the first, with just one different step. Rather than freely diffusing throughout the cell, reduced PCN or PCA intercalate with eDNA and transfer electrons onto the eDNA complex.â¹ Electrons are transferred through stacked pi-orbitals of DNA until they reach intercalated PYO, which then acquires electrons and uncouples from the eDNA double helix.â¹ Both methods of PMEET have been shown to be energetically favorable, fast and efficient.â¹ Individual experiments suggest that both methods likely occur endogenously and simultaneously.â¹ Additionally, this study was the first time eDNA has been identified as not just a major structural component of biofilms, but also as an important component of biofilm metabolism.â¹
BIOTECHNOLOGICAL AND BIOPHARMACEUTICAL APPLICATIONS OF SMEES
Small molecule extracellular electron shuttles are used in several biotechnological and biopharmaceutical applications. One of the most important applications of SMEES is their use in microbial fuel cells. Microbial fuel cells (MFCs) are essentially batteries made from bacterial cultures in which the electrochemical energy produced by bacterial metabolism is harnessed to power electrical devices.¹ MFC current generation has been shown to benefit from both the endogenous production and exogenous addition of SMEES.¹ MFCs represent an exciting form of sustainable energy as energy production does not rely on external stimuli. However, they are currently hampered by high electrical resistances which constrain electrical outputs.
Apart from microbial fuel cells, SMEES have found use in the bioremediation of toxic heavy metals and crude oil.¹ They have also been used as agricultural probiotics, stimulating growth of beneficial agricultural microbial symbionts and suppressing harmful ones.¹â· Furthermore, some pharmaceuticals contain chemical moieties analogous to those of known SMEES.¹â¸
Redox-active natural products are a diverse group of organic molecules produced by a wide range of bacteria. Many redox active natural products may function as electron shuttles. Future research, namely environmental metabolomics and metagenomics, could potentially reveal that small molecule extracellular electron transfer is a widespread strategy for survival by indirect aerobic respiration.
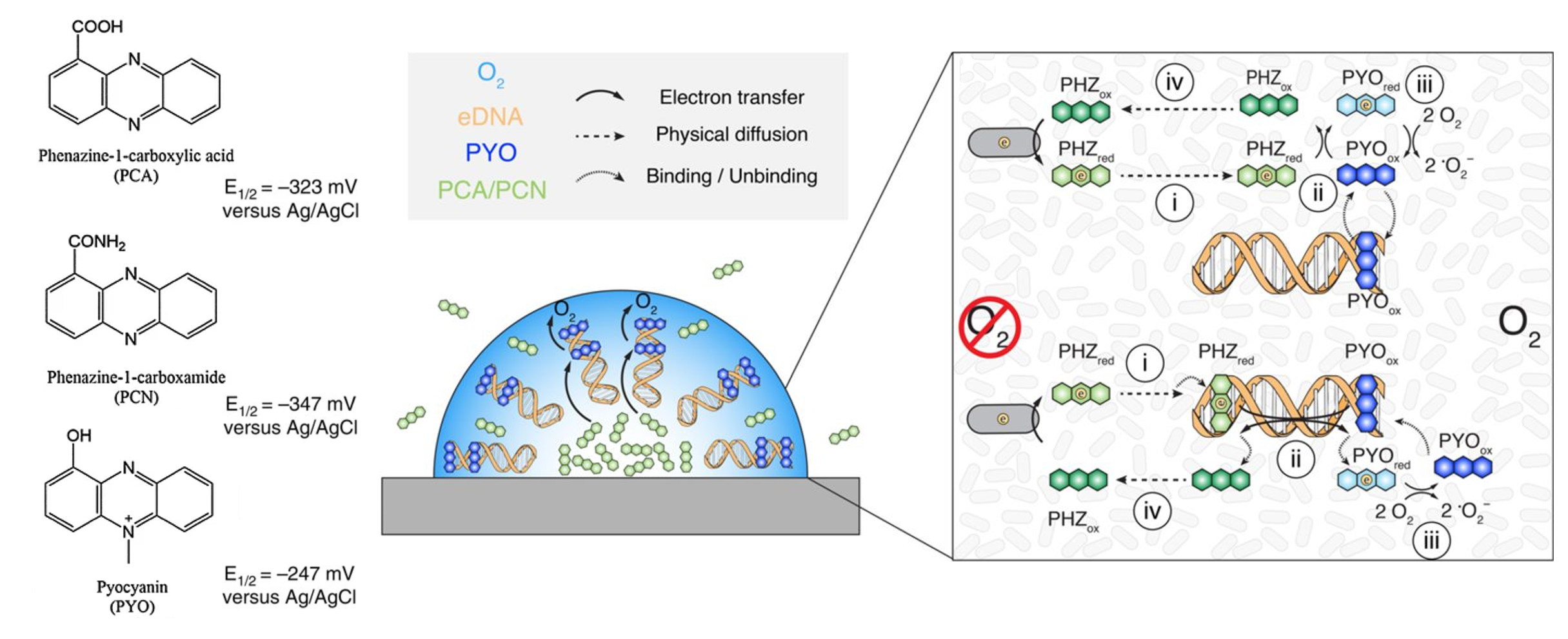
(A) Structures and redox potentials of the electron shuttles PCA, PCN, and PYO.
(B) Organization of several phenazines, eDNA, and oxygen concentration in a Pseudomonas
aeruginosa biofilm.
(C) Two proposed mechanisms for phenazine mediated extracellular electron transfer. The top without DNA charge
transfer and the bottom with. Steps are explained in text. PHZ indicates a lower redox phenazine, either PCA or PCN. Saunders (2020) Cellâ¹
REFERENCES
- Anraku, Y. Bacterial electron transport chains. Annual review of biochemistry, 57(1), 101-132. (1988).
- Morris, R. L., & Schmidt, T. M. Shallow breathing: bacterial life at low O2. Nature Reviews Microbiology, 11(3), 205-212. (2013).
- Schauer, R., et. al. Succession of cable bacteria and electric currents in marine sediment. The ISME journal, 8(6), 1314-1322. (2014).
- Steidl, R. J., Lampa-Pastirk, S., & Reguera, G. Mechanistic stratification in electroactive biofilms of Geobacter sulfurreducens mediated by pilus nanowires. Nature Communications, 7, 12217. (2016).
- Shi, L., et. al. Extracellular electron transfer mechanisms between microorganisms and minerals. Nature Reviews Microbiology, 14(10), 651-662. (2016).
- Schmidt, R., et. al. Microbedriven chemical ecology: past, present and future. The ISME journal, 13(11), 2656-2663. (2019).
- Pelaez, F. The historical delivery of antibiotics from microbial natural products– can history repeat?. Biochemical pharmacology, 71(7), 981-990. (2006).
- Bilal, M., et. al. Engineering Pseudomonas for phenazine biosynthesis, regulation, and biotechnological applications: a review. World Journal of Microbiology and Biotechnology, 33(10), 191. (2017).
- Saunders, S. H., et. al. Extracellular DNA promotes efficient extracellular electron transfer by pyocyanin in Pseudomonas aeruginosa biofilms. Cell, 182(4), 919-932. (2020).
- Price-Whelan, A., Dietrich, L. E., & Newman, D. K. Rethinking Secondary Metabolism: physiological roles for phenazine antibiotics. Nature chemical biology, 2(2), 71-78. (2006).
- Friedheim, E. A. Pyocyanine, an accessory respiratory enzyme. The Journal of experimental medicine, 54(2), 207. (1931).
- Ramos, I., Dietrich, L. E., Price-Whelan, A., & Newman, D. K. Phenazines affect biofilm formation by Pseudomonas aeruginosa in similar ways at various scales. Research in microbiology, 161(3), 187-191. (2010).
- Bellin, D. L., et. al. Electrochemical camera chip for simultaneous imaging of multiple metabolites in biofilms. Nature communications, 7(1), 1-10. (2016).
- Rabaey, K., Boon, N., Höfte, M., & Verstraete, W. Microbial phenazine production enhances electron transfer in biofuel cells. Environmental science & technology, 39(9), 3401-3408. (2005).
- Dasgupta, D., Kumar, A., Mukhopadhyay, B., & Sengupta, T. K. Isolation of phenazine 1, 6-di-carboxylic acid from Pseudomonas aeruginosa strain HRW. 1-S3 and its role in biofilm-mediated crude oil degradation and cytotoxicity against bacterial and cancer cells. Applied microbiology and biotechnology, 99(20), 8653-8665. (2015).
- Dahlstrom, K. M., McRose, D. L., & Newman, D. K. Keystone metabolites of crop rhizosphere microbiomes. Current Biology, 30(19), R1131-R1137. (2020).
- Borrero, N. V., et. al. Phenazine antibiotic inspired discovery of potent bromophenazine antibacterial agents against Staphylococcus aureus and Staphylococcus epidermidis. Organic & Biomolecular Chemistry, 12(6), 881-886. (2014).
- Singh, S., et al. “In vitro evaluation of a new drug combination against clinical isolates belonging to the Mycobacterium abscessus complex.” Clinical Microbiology and Infection 20.12 (2014): O1124-O1127.
WRITTEN BY FISHER PRICE
M.S. Biology, UC San Diego 2021, Jensen Lab, Center for Marine Biotechnology and Biomedicine, Scripps Institution of Oceanography
FROM SALTMAN QUARTERLY VOL. 18
To read the original version, please click here. To read the full version on our website, please click here. To read more individual articles, please click here.