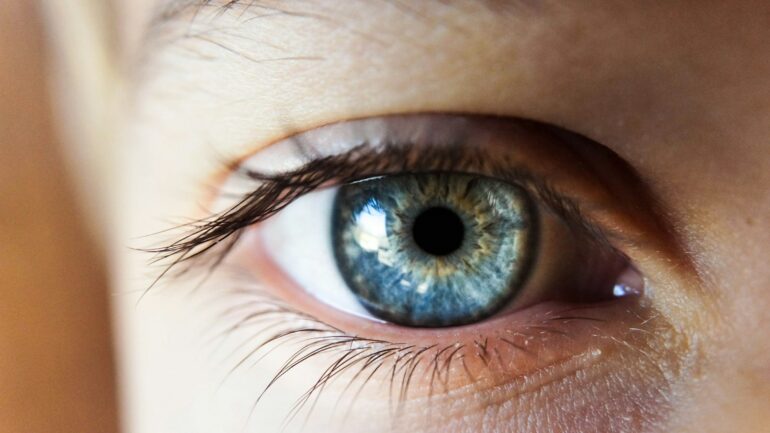
What do you think makes you physically identifiable? Is it your height? Or maybe your unique eye color? All of these visible traits make up your phenotype. That is only one definition of phenotype, however. Phenotype can also mean the physical symptoms of an ailment. When you go to the doctor, how are you diagnosed with an illness or a disease? Usually, the doctor will search for detectable symptoms like a runny nose or a cough—your phenotype—and then diagnose and prescribe medication based on them. While this phenotype-first method is accurate enough to diagnose simple things like a cold or flu, it is oftentimes not precise enough to identify more complicated ailments like genetic disorders. In the United States, 7.4 million patients are misdiagnosed every year, which may result in patients taking the wrong medication, serious harm, or even death. There might be a way to improve these odds, however, by looking at the human genome.
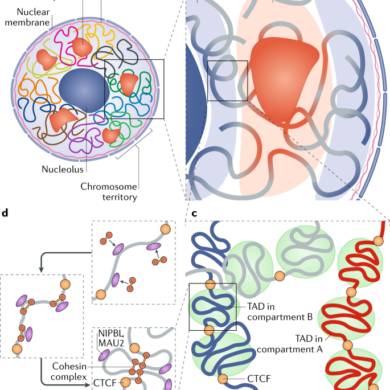
Our genes carry DNA, which codes for the proteins that give us our physical traits. For example, a person who has dark brown hair does so because a portion of their DNA (a gene) encodes for that trait via nucleotide bases (adenine, thymine, guanine, and cytosine) that carry genetic information. Through the cellular processes of transcription and translation, that dark brown hair gene synthesizes a protein that carries out functions in the body to make that phenotype a reality. Frequently, several genes are involved in producing the proteins required for a trait. All the genes in your body make up your entire genome. A mutation in one or several genes (such as an incorrectly placed nucleotide, the deletion of a nucleotide of a gene, or the presence of extra copies of a gene) can manifest into genetic disorders.

Double-stranded DNA (Source)
Down syndrome is a genetic disorder that is diagnosed using the phenotype-first approach after birth. First, doctors identify some of the common physical signs of Down syndrome in potential patients: short neck, flattened face, decreased muscle tension, etc. To confirm the diagnosis, genetic testing will be done to see whether the patient has an extra copy of chromosome 21—the genetic cause of Down syndrome. While Down syndrome is highly understood and has very distinct physical traits, this is not the case for genetic disorders like Huntington’s Disease, whose early symptoms—like difficulty concentrating and irritability—are subtle and often mimic other disorders. Thus, the phenotype-first approach is not the most viable in these more complex cases where the phenotypic traits of an illness are not clear or distinct. Excitingly, recent developments in the field of genetic research have outlined a potential solution to this issue: the genotype-first approach. Unlike the phenotype-first approach, the genotype-first approach starts in the human genome. Instead of identifying clinical symptoms and then trying to find the genetic changes that may explain those symptoms, researchers study the physical traits of patients with specific gene variants, which essentially is the opposite of the phenotype-first approach. Two studies demonstrate the benefits of this application.
STUDY 1: TPSAB1 GENE
Jonathan J. Lyons of the National Institute of Allergy and Infectious Diseases conducted the first study using this genotype-first approach. Lyons and his colleagues connected gastrointestinal issues like irritable bowel syndrome to elevated tryptase levels in the body (tryptase is a protein found in white blood cells that breaks down other proteins) via a hypothesized genetic factor. The TPSAB1 gene was high on their list of potential causes as it specifically encodes for tryptase. To test this, they genetically identified people with variations in the TPSAB1 gene—particularly if they had extra copies of it—through genotyping. Genotyping is a common laboratory test used to determine the DNA sequence of specific areas of an individual’s genome, and one’s genotype, or unique DNA sequence. Essentially, your genotype explains why you look the way you do your phenotype. Comparison of one’s genotype to a reference sample can help identify any genetic variations present. Genotyping can also determine the number of copies of a particular gene, otherwise known as copy number variants, present in one’s genome. Because DNA is made up of nucleotide bases that pair together (adenine and thymine bond together while guanine and cytosine do the same), one sequence can stick to other nucleotide sequences with the proper matching bases. So, Lyon used complementary DNA strands that stuck to the TPSAB1 gene, thus allowing him to identify how many copies were present in each individual. When compared to the reference sequences, the patients with duplications of the TPSAB1 gene were found to not only have elevated tryptase levels but also gastrointestinal issues. Thus, the genotype-first approach allowed for the establishment of new relationships between gene variants and the clinical symptoms of gastrointestinal ailments that were previously unknown, which will largely help in diagnosing patients moving forward. More studies like this are needed to develop a knowledge of gene variations and how they manifest into physical symptoms.
STUDY 2: ACSF3 GENE
The second study was performed by Jennifer L. Sloan, M.D., of the Genetics and Molecular Biology Branch of the National Human Genome Research Institute in the United States. Sloan identified that mutations like deletions in the ACSF3 gene were the cause of a rare metabolic disorder known as Combined Malonic and Methylmalonic Aciduria (CMAMMA). She identified these mutations by sequencing the protein-coding portions of patient DNA via a widely used method named Sanger sequencing, in which DNA complementary to the strand of interest is synthesized in fragments of various sizes. These fragments are separated from shortest to longest, with the terminating base (adenine, thymine, guanine, or cytosine) being fluorescently labeled. The identity of each terminating base is recorded from the shortest length fragment to the longest one, which gives the final DNA sequence. Several of the patients with these genetic variations had severe neurological symptoms like memory loss and seizures as a result of their CMAMMA except one sixty-six-year-old woman who had seemingly perfect health. However, when her DNA was sequenced, it was uncovered that she had several CMAMMA-causing ACSF3 gene mutations. Further testing using MRI readings and other examinations revealed that she did have CMAMMA but suffered from much subtler symptoms due to getting the disease at an older age. For example, her memory issues did not match CMAMMA’s original diagnostic criteria. Without the genotype-first approach, Dr. Sloan may have never discovered these milder symptoms of CMAMMA that can now be used for patient diagnosis. Further studies could uncover many more physical manifestations of ailments that we are not yet aware of, especially in the cases of early or late-onset diseases where symptoms and their intensities vary greatly.

DNA genotyping and sequencing (Source)
LIMITATIONS, BENEFITS, AND THE FUTURE:
Although the genotype-first approach offers several advantages to the field of diagnostic medicine, it does have several limitations. At the moment, there is a glaring lack of diversity in genomic studies. So far, genetics research has mostly been done in white populations, which leads to biased results because genetics-related diseases do not manifest the same in all populations. These studies must include diverse, multi-racial, and ethnic populations to be as all-encompassing as possible. Furthermore, factors like age should be considered as well. Efforts have been made for genomic studies to be more inclusive: Genentech, a biotechnology company that creates new medicines for severe sicknesses, is a great example. They are actively working to expand the populations of their clinical trials to include underrepresented groups so that they can receive the healthcare best suited for them and their genetics. Overall, much more research is needed before the genotype-first approach becomes a truly viable method of diagnosis for all people.
The most obvious benefit of the genotype-first approach is that it will hopefully lower the patient misdiagnosis rate, but it’s also important to acknowledge that this method brings us one step closer to the future of medicine: preventative medicine. Currently, with the phenotype-first approach, medicine is reactive. Only when physical symptoms of an ailment manifest, which could take a long time, can doctors prescribe medication to treat the illness. But, with the genotype-first approach, patients can be screened for genomic variations associated with certain diseases before there are any signs of physical illness. If they have these genomic mutations, steps can be taken to either reduce their risk of developing the illness in the future or to treat the disease at its early stages, improving their chances of a healthy recovery.
Sources:
- https://www.genome.gov/genetics-glossary/Phenotype
- https://effectivehealthcare.ahrq.gov/products/diagnostic-errors-emergency-updated/research#field_report_title_1
- https://www.genome.gov/For-Patients-and-Families/Genetic-Disorders
- https://www.nichd.nih.gov/health/topics/down/conditioninfo/diagnosis
- https://www.nichd.nih.gov/health/topics/down/conditioninfo/symptoms
- https://www.aamc.org/news/rare-diseases-difficult-diagnose-cures-hard-come
- https://www.huntingtonswa.org.au/information/living-with-hd/early-stages
- https://www.cell.com/ajhg/fulltext/S0002-9297(22)00538-9#secsectitle0010
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5397297/
- https://www.testing.com/tests/tryptase/
- https://www.idtdna.com/pages/applications/genotyping
- https://www.genome.gov/genetics-glossary/Copy-Number-Variation
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3163731/
- https://www.genomicseducation.hee.nhs.uk/genotes/knowledge-hub/sanger-sequencing/
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9892776/
- https://www.fredhutch.org/en/news/center-news/2019/06/lack-diversity-genetic-research-problem.html
- https://www.gene.com/patients/clinical-trials/advancing-inclusive-research
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1299205/